sklearn.linear model.ElasticNet
Jump to navigation
Jump to search
A sklearn.linear_model.ElasticNet is an ElasticNet System within sklearn.linear_model class.
- AKA: ElasticNet, linear model.ElasticNet.
- Context
- Usage:
- 1) Import ElasticNet Regression model from scikit-learn :
from sklearn.linear_model import ElasticNet - 2) Create design matrix
Xand response vectorY - 3) Create ElasticNet object:
ENreg=ElasticNet([n_iter=300, tol=0.001, alpha_1=1e-06, alpha_2=1e-06, lambda_1=1e-06, lambda_2=1e-06, compute_score=False, fit_intercept=True, normalize=False, copy_X=True, verbose=False]) - 4) Choose method(s):
- Fit the ElasticNet model with coordinate descent to the dataset:
ENreg.fit(X, Y[, check_input])) - Predict Y using the linear model with estimated coefficients:
Y_pred = ENEreg.predict(X) - Return coefficient of determination (R^2) of the prediction:
ENreg.score(X,Y[, sample_weight=w]) - Compute elastic net path with coordinate descent:
ENreg.path(X, y[, l1_ratio, eps, n_alphas,...]) - Get estimator parameters:
ENreg.get_params([deep]) - Set estimator parameters:
ENreg.set_params(**params)
- Fit the ElasticNet model with coordinate descent to the dataset:
- 1) Import ElasticNet Regression model from scikit-learn :
- Example(s):
| Input: | Output: |
#Importing modules
#Calculaton of RMSE and Explained Variances
# Printing Results
#plotting real vs predicted data
|
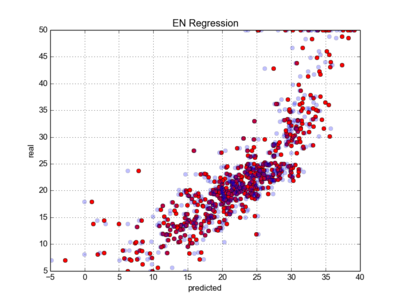
|
- Counter-Example(s):
- See: Regression System, Regularization Task, Ridge Regression Task, Bayesian Analysis.
References
2017
- http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.ElasticNet.html
- QUOTE:
class sklearn.linear_model.ElasticNet(alpha=1.0, l1_ratio=0.5, fit_intercept=True, normalize=False, precompute=False, max_iter=1000, copy_X=True, tol=0.0001, warm_start=False, positive=False, random_state=None, selection=’cyclic’)
- QUOTE:
- Linear regression with combined L1 and L2 priors as regularizer.
- Minimizes the objective function:
1 / (2 * n_samples) * ||y - Xw||^2_2 + alpha * l1_ratio * ||w||_1 + 0.5 * alpha * (1 - l1_ratio) * ||w||^2_2
- If you are interested in controlling the L1 and L2 penalty separately, keep in mind that this is equivalent to:
a * L1 + b * L2- where:
alpha = a + b and l1_ratio = a / (a + b)- The parameter l1_ratio corresponds to alpha in the glmnet R package while alpha corresponds to the lambda parameter in glmnet. Specifically, l1_ratio = 1 is the lasso penalty. Currently, l1_ratio <= 0.01 is not reliable, unless you supply your own sequence of alpha.